Making the Connection
- Published1 Aug 2016
- Reviewed1 Aug 2016
- Author Susan Gaidos
- Source BrainFacts/SfN
For six painstaking years, Jeff Lichtman and his team at Harvard University worked to assemble a complete 3-D map of a bit of brain so tiny it is about the breadth of a human hair. Driving Lichtman’s efforts and those of others trying to sort out how the brain wires itself together is the fact that somehow consciousness, personality, and memories arise from all those cells and connections. “That’s what we’re seeking,” he says.
Finished in 2015, that comprehensive reconstruction represented the largest portion of mammalian brain ever rendered in full detail – it measured 1,500 cubic microns and documented every cell type and connection. Yet that small sample illustrates the enormous complexity of the brain as it contained fragments of 1,600 neurons and 1,700 synapses. What’s more, it revealed a fundamental truth about neurons and the brain: forming synapses requires more than two neurons being close to one another. What prompts the synaptic connection remains to be discovered.
Developing wiring diagrams ranging in scale from the individual synapse like Lichtman’s to the whole brain is a high priority for the BRAIN Initiative – a research effort launched by President Barack Obama in April 2013 to catalyze and speed advances in neuroscience by deploying cutting-edge computer science, physics, biology, and chemistry to develop transformative new tools. Understanding how the brain processes information at the level of the circuit in addition to the physical structure of the connections could provide insights into how our brains behave when they are healthy, and what goes wrong in disorders like autism and schizophrenia when those circuits fail to hook up properly.
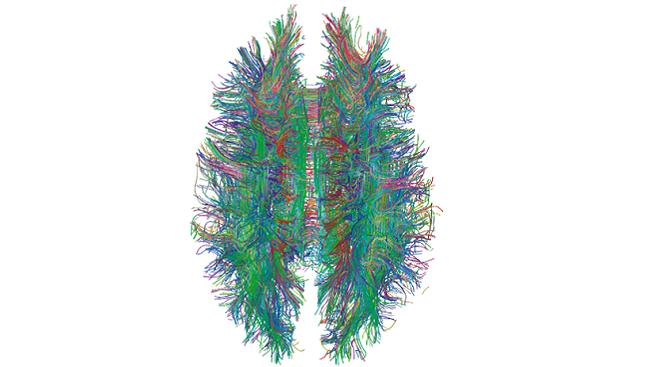
The effort to create a catalog of all the connections between the brain’s cells – often called a connectome – is daunting. The human brain is the most complex biological structure in the known universe. It houses 86 billion neurons connecting to each other through 150 trillion synapses over thousands of miles of axons. In the past decade or so scientists have started to develop a new set of tools needed to rapidly enhance the field’s ability to unravel the brain’s mysteries.
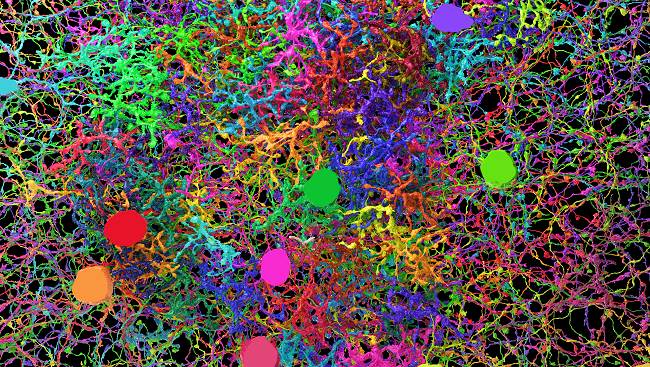
In order to help researchers discriminate between neurons, Lichtman and Joshua Sanes, also at Harvard, developed a method called Brainbow to color neurons with fluorescent dye. Basically, Lichtman and Sanes engineered mice with a gene encoding four different fluorescent proteins. That gene is susceptible to random cutting and as a result random mixtures of colors populate each neuron. In that way, nearly 90 colors are available.
While the technique helped researchers identify individual neurons, the resolution wasn’t adequate to explore complex circuits. In producing the detailed 3-D reconstruction of the mouse neocortex, Lichtman needed new tools. His group built a machine to automate the cutting process, slicing the speck of brain into wispy thin slivers, measuring 30 nanometers, or 1/1,000 the thickness of a human hair. By mounting the slices on a tape, similar to a filmstrip, the scientists delivered samples in a conveyor-belt fashion to a specialized electron microscope. The team then aligned the resulting images and employed a combination of software and human input to trace the long tendrils of axons and other parts of cells through the images.
“We know that the brain is made up of nerve cells that are connected together, but no one had ever been able look at a piece of brain and see how many wires are there, or how many synapses are there or how often the same axon contacts the same dendrite at different sites,” Lichtman says. “Previous imaging techniques were just too limiting.”
Lichtman will use that same process combined with an even more powerful microscope in a project sponsored by the Intelligence Advanced Research Projects Activity (IARPA) aimed at quantifying why brains are so good at learning in an effort to improve the algorithms for machine learning and artificial intelligence. Neural network algorithms are already sophisticated enough to beat humans at the complex Chinese strategy game Go, and are learning how to drive automobiles, but computers still can’t match human intelligence in more complex settings such as understanding language.
Even though, computers recognize objects extremely well, in especially difficult cases, humans do it better. H. Sebastian Seung, of Princeton University, is taking advantage of that human facility. Faced with the realization that a single person trying to trace the circuits in a cubic millimeter of human cortex would need nearly million years to complete the task, Seung decided to engage citizen scientists to help map the mouse retina. While still at the Massachusetts Institute of Technology, Seung’s graduate student Mark Richard developed Eyewire, an online computer game funded by in part by the National Institutes of Health.
Eyewire gamers color in the branches of neurons as they move through a 3-D electron microscope image of a section of mouse retina. Players earn points, participate in community building events, and even become authors on papers. In 2014, Seung published a paper detailing a wiring diagram for a circuit including starburst amacrine cells – a type of neuron with dendrites extending in all directions that has been implicated in detecting moving objects. The wiring diagram indicates the circuit feeding into the amacrine cells operates on a time delay to allow the retina to detect motion and direction.
From Cells to Circuits
Understanding how small networks of neurons work together to make sense of visual information serves as a starting point for piecing together the rules, or regularities of visual processing in the brain. To describe the principles linking circuits to a particular function, scientists must determine how networks are built.
The Allen Institute for Brain Science’s R. Clay Reid set out to do just that. In March 2016, he announced the completion of an effort to link the wiring of a tiny bit of a mouse visual cortex to its function. The work began a decade prior to the announcement and was the largest network at the time of its announcement to detail the connections between neurons in the visual cortex. The network identified 50 neurons and 990 synapses.
In order to do so, the researchers genetically engineered a mouse’s neurons to fluoresce whenever the calcium levels increased around the neuron – a measure of activity. By peering through a skull window, the team could record the neurons firing while the animal watched a video. A second part of the experiment had the researchers slicing that brain into thousands of layers less than 50 nanometers thick. The researchers then made electron microscope images of each layer and began mapping out the connections between those neurons. The effort revealed the first structural evidence supporting the idea that neurons performing similar tasks are more likely to be connected than those performing different tasks.
“Part of what makes this study unique is the combination of functional imaging and detailed microscopy,” Reid told the Allen Institute. “The microscopic data is of unprecedented scale and detail.”
Reid will use this combination of methods to tackle IARPA’s challenge to reverse-engineer one cubic millimeter of the mouse brain by recording the activity and connectivity of 100,000 neurons of the animal that had completed visual perception and learning tasks. Reid is part of a team of researchers including Princeton’s Seung and others from the Allen Institute and Baylor College of Medicine. In addition to Reid’s team, IARPA has engaged two other teams in its Machine Intelligence from Cortical Networks (MICrONS) project as well. Lichtman will participate with a Harvard University team led by David Cox. Another team includes researchers from Carnegie Mellon University and the Wyss Institute for Biologically Inspired Engineering at Harvard University.
The IARPA project aims to improve artificial intelligence one day. However, Lichtman notes the information generated will also provide valuable insights into how our brains make us human as scientists seek “those regularities that will allow us to understand how … information about the world gets physically built into your brain to create memories, personality traits and skills that you carry with you throughout your life.”
This article was produced in conjunction with the Brain Initiative Alliance, a supporting partner of BrainFacts.org.
CONTENT PROVIDED BY
BrainFacts/SfN
References
Kasthuri N, Hayworth KJ, Berger DR, Schalek RL, Conchello JA, et al. Saturated Reconstruction of a Volume of Neocortex. Cell. 162(3):648-61 (2015).
Kim JS, Greene MJ, Denk W, Seung HS; EyeWirers, et al. Space-time wiring specificity supports direction selectivity in the retina. Nature. 509(7500):331-6 (2014).
Lee WC, Bonin V, Reed M, Graham BJ, Hood G, Glattfelder K, Reid RC. Anatomy and function of an excitatory network in the visual cortex. Nature. 532(7599):370-4 (2016).
Livet J, Weissman TA, Kang H, Draft RW, Lu J, Bennis RA, Sanes JR, Lichtman JW. Transgenic strategies for combinatorial expression of fluorescent proteins in the nervous system. Nature. 450(6293):56-62 (2007).
Mikula S, Denk W. High-resolution whole-brain staining for electron microscopic circuit reconstruction. Nature Methods. 12(6):541-6 (2015)
Morgan JL, Berger DR, Wetzel AW, Lichtman JW. The Fuzzy Logic of Network Connectivity in Mouse Visual Thalamus. Cell. 165(1):192-206 (2016).